PepCNN deep learning tool for predicting peptide binding residues in proteins using sequence, structural, and language model features | Scientific Reports

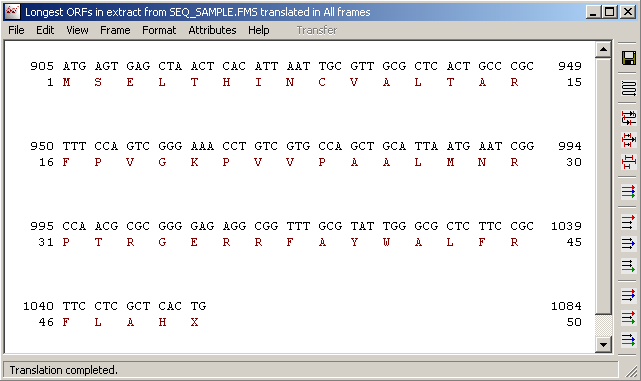
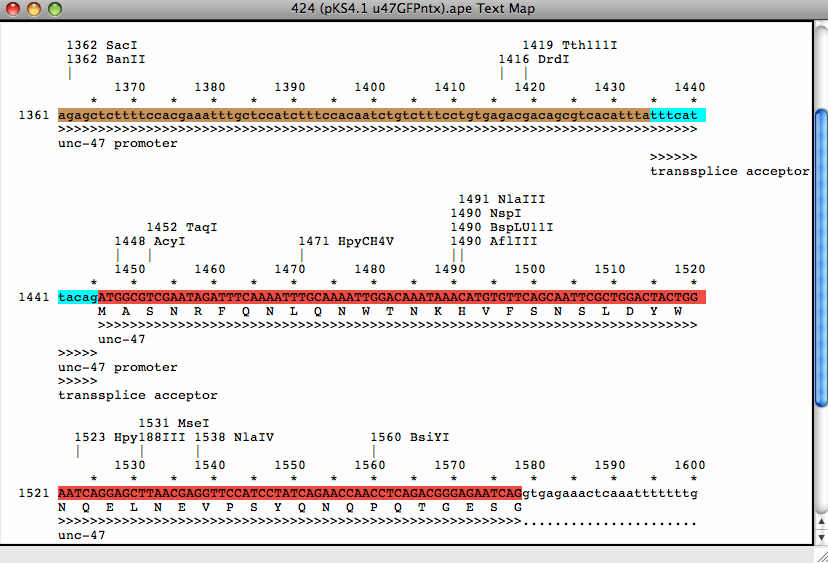
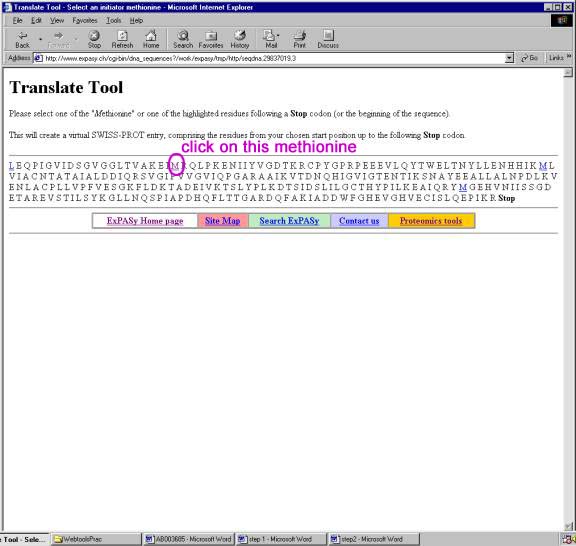
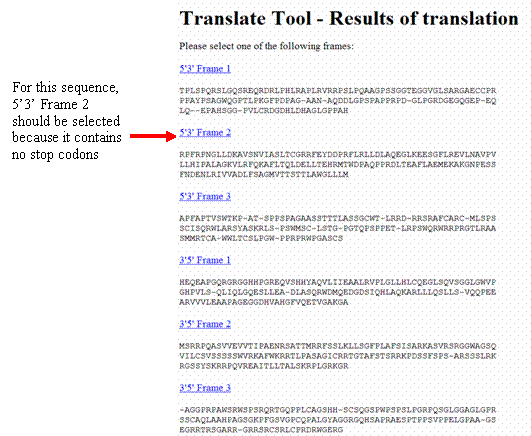
In silico translation of truncated sequences in the ExPASy Translate... | Download Scientific Diagram

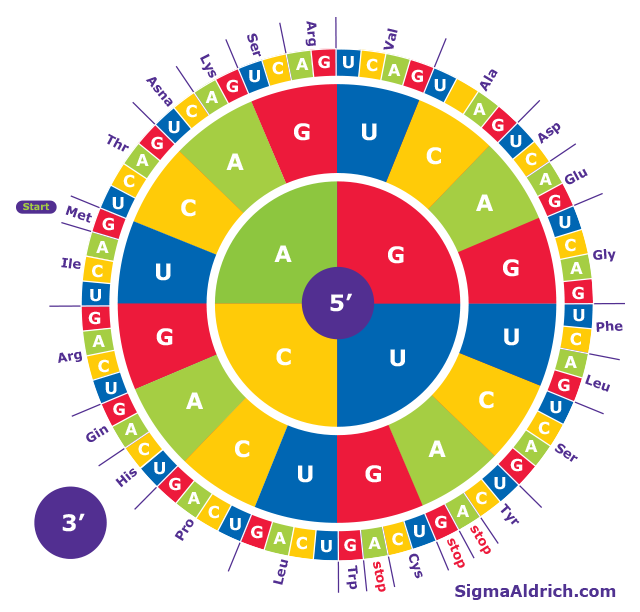
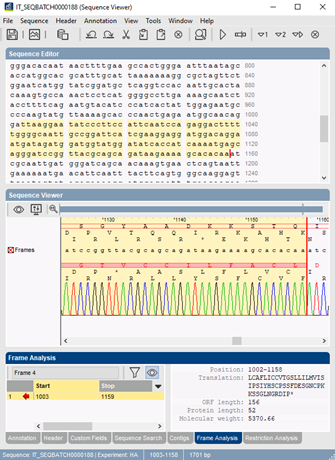
ExPASy | Translate a nucleotide sequence and select the correct reading frame of the polypeptide - YouTube
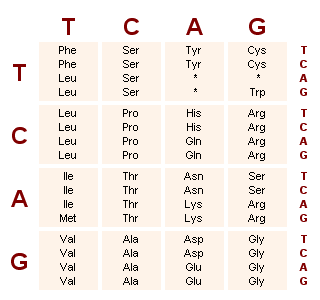
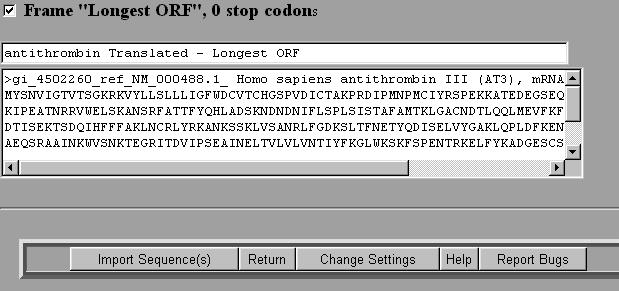
![PDF] Virtual Ribosome—a comprehensive DNA translation tool with support for integration of sequence feature annotation | Semantic Scholar PDF] Virtual Ribosome—a comprehensive DNA translation tool with support for integration of sequence feature annotation | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/5b35b31050c9cf2242b96df3e3aa6b5856169564/4-Figure2-1.png)